Multiomics technologies for infectious disease research and the role of dietary iron in promoting pneumococcal diseases
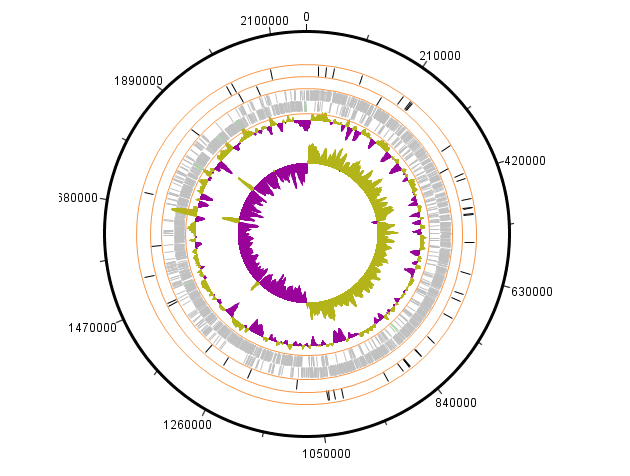
The uptake of micronutrients, such as iron in the human bloodstream and mucosal surfaces, is an essential requirement for microbes to cause diseases. Bacteria are often in “battle” with their human host over maximizing the use of iron for a variety of cellular processes linked to pathogenesis such as biofilm formation and toxin expression (). In a series of research reports, I used advanced multiomics technologies such as proteomics, non-coding, and mRNA expression and genome arrays to establish the important role of iron in pneumococcal diseases. These were some of the first studies describing the use of high throughput technologies in microbiology research.
Peer-Reviewed Publications
-
Identification of novel non-coding small RNAs from Streptococcus pneumoniae TIGR4 using high-resolution genome tiling arrays. 2010. [Abstract] [Full Paper]
Pratik Shah†, Kumar R†, Swiatlo E, Burgess SC, Lawrence ML, Nanduri B
(†Contributed equally)
BMC Genomics. PMID: 20525227
-
Differential gene expression in Streptococcus pneumoniae in response to various iron sources. 2009. [Abstract] [Full Paper]
Radha Gupta, Shah P, Swiatlo E
Microbial Pathogenesis. PMID: 19464356
-
Quantitative analysis of Streptococcus pneumoniae TIGR4 response to in vitro iron restriction by 2-D LC ESI MS/MS. 2008. [Abstract] [Full Paper]
Bindu Nanduri, Shah P, Ramkumar M, Allen EB, Swiatlo E, Burgess SC, Lawrence ML
Proteomics. PMID: 18491321